ALERT! Warning to America’s Savviest Investors, Savers and Retirees:
Inflation Rocket Train “Accelerating Fast”
Painful Carter-Era, 70s-Style “500% Inflation” Looming
- Will Your Savings Survive Hyperinflation?
- What Happens When the Banks Get Wiped Out?
- What If Taxes Soar Not Only for the “Rich?”
- Can You Survive If Stocks Collapse?
Dear Valued Reader:
In less than a decade of reckless government expansion, the fundamentals of successful investing, planning and living have undergone a dramatic and potentially dangerous transformation. The ground has literally changed under your feet!
The number of complex legal, regulatory, medical and retiree-unfriendly tax traps has mushroomed to the point that ignorance is downright dangerous. And with President Biden’s newest multi-trillion dollar “social safety net” tax hike, it’s about to get worse. The upshot is cash-hungry bureaucrats are now looking for full control over everything from cash in the bank to private retirement assets held anywhere they can get their hands on it. And they want to suppress alternative asset protection and wealth building information that can make people more self-reliant — like the kind in this letter.
To prosper in Biden’s America (yes, it’s still possible and I’ll show you how), you need to consider some new developments and plan accordingly. The information I want to send you is essential to anyone who wants to protect their wealth, prepare for their retirement or is already in retirement.
I am convinced that the U.S. is now facing some of the most serious economic and political threats in its history. After nearly 40 years of unprecedented prosperity, the U.S. economy is now in shambles.
Millions of Americans have lost their jobs… their life savings… and soon it will be their homes. While the U.S. military remains the most powerful force in the world, the most immediate threats America now faces are closer to home.
These include: Companies forced out of business… soaring government spending that will lead to hyperinflation… runaway taxes… older citizens no longer able to retire and live what used to be known as “their golden years.” And more…
Fortunately, you DON’T have to just sit by and do nothing. There are simple, easy, inexpensive steps you can take right now to protect your family. BUT…
If You Can’t Answer “YES” to These Questions,
You May Be Behind the “8‑Ball.”
Have you packed away every penny you legally can for your retirement? If cash-hungry Federal bureaucrats dilute the value of IRAs and other retirement accounts, would you be affected? What is your exposure to and what preparations have you made for the coming housing bubble collapse?
You deserve to be just as protected against the coming inflation wave as any tech giant or financial titan of Wall Street. That’s why I want to send you the inside information and strategies they already know, before it’s too late.
As I am about to show you with shocking urgency, now is the time to make basic preparations to come out ahead in the collapsing value of dollar-denominated assets — to not be a victim or a sucker. I have no intention of being either, and I know you feel the same way.
At the end of this letter, I will tell you how to get the newest, most authoritative source of information about hidden inflation, how it is ripping you off and what you can do to protect yourself. The Ultimate Hyperinflation Defense Manual is probably among the most important guides I have ever offered. You can’t find this information anywhere else and you haven’t a moment to lose.
A Dire Warning from TWO Past Federal Reserve Board Chairs!

As the editor of The Bob Livingston Letter®, I pay very close attention to recently resigned or soon-to-be-departed high-level U.S. officials (once they are out of office many are remarkably candid — even as the “mainstream” media stop reporting their words). In my experience, recently-retired officials often hint at dangers to come, leaving a trail of “breadcrumbs” in which Americans like you and I should pay close attention.
That’s why it’s so important to note the departing warning of former Federal Reserve Board Chairman Alan Greenspan (a man noted for understatement) — who flatly told Congress “serious economic disruptions are in the works” due to trillions of dollars in Federal obligations that will be visited upon taxpayers and retirees like a gigantic balloon payment. And that was well before the trillions in helicopter money started raining down on America.
Now listen to Former Fed Chair Paul Volker who presided over 70’s-era inflation and was even more blunt. He warned that the chances of an all-out dollar collapse due to runaway spending were greater than 50-50.
These former insiders are referring to a level of unfunded liability by the Federal government which is so gigantic as to boggle the mind. To give you some idea of the scope of this problem, the government would have to extract $473,000 from each and every U.S. household to cover the rapidly approaching national balloon payment on unfunded liabilities!
Obviously, something is going to have to “give.” This is why I want to rush you information on how Treasury bureaucrats are forming emergency “contingency” plans to freeze IRA and 401(k) accounts in the event of a national financial panic — similar to the “banking holidays” used by the Franklin Roosevelt administration to bilk millions of Americans during the Great Depression.
You especially need this expanded, updated and revised manual because “passive” retirement planning is a thing of the past. To get the most out of your life over 50, you can no longer listen to only one or two paid “experts,” make your house payments and “assume” the best. It’s not enough anymore, not by a long shot! This is true if you’re 10-15 years away from retirement and particularly if you are already in retirement. (The latter group has even more to lose through inaction.)
How to Live Through Hyperinflation.
In the past few months, prices have been skyrocketing. Lumber prices have almost tripled. Real estate is soaring. Prices for some commodities have shot through the roof. And let’s not even talk about the grocery store… but don’t be fooled! The current, temporary period of inflation will soon be followed by even more massive inflation. That’s because President Biden and his allies in Congress have embarked on the biggest government spending spree in history. This socialistic expansion of government will inevitably lead to a devaluation of the U.S. dollar (hyperinflation) that could wipe out your life savings.
Imagine a world of $20 per gallon of gasoline… $50 hamburgers… and average housing rentals costing $10,000 a month or more. The economic stimulus packages of the last few years alone will amount to $6 trillion… nearly 7 times what the Troubled Assets Relief Program (TARP) program spent! And the stimulus plan is only the tip of the iceberg!
When you add in the unfunded liabilities, that’s trillions the government doesn’t have over the next few years. Whether the government borrows it or prints it, an increase that big in the money supply is going to do only one thing: Send inflation through the roof! The last time the U.S. government tried to spend its way out of a recession (in the 1970s) the inflation rate increased by 500%!
The Ultimate Hyperinflation Defense Manual is probably among the most important guides I have ever offered. You can’t find this information anywhere else and you have my personal word of honor you’ll be glad you accepted my special offer below. Why? It will bring you up to speed on:
- Shockingly deceptive government statistics that will make your blood boil — on a scale unprecedented in our history. All designed to encourage millions of uninformed U.S. investors to keep their money in highly-vulnerable dollar-denominated financial instruments only. Even when everyone in the know is running for the exits.
- Virtually unreported dire warnings from America’s top investors and former top government officials. Even the comptroller of the U.S. confirms what investors and steadily-rising gold prices are pointing to: The impossible financial pickle the government finds itself in.
- Understand all the NEW options you have to protect yourself from the coming inflation tsunami — far above and beyond what smart people had in the 1970s to make themselves rich even as most Americans saw their life savings ravaged by inflation.
- The government’s secret manipulation of the gold market downward — and why hiding grim inflation statistics from millions of investors is coming to a nasty and brutish end.
Hard Times Ahead if You Are on a Fixed Income… or Just Unprepared.
Accelerating inflation is a threat to your future comfort, big time! It means your stock portfolio is worth less than you think or need. It also reduces your future ability to pay for fuel, electricity, food, healthcare and other basic daily needs. And it is especially merciless to people who assume they can live comfortably on a fixed income.
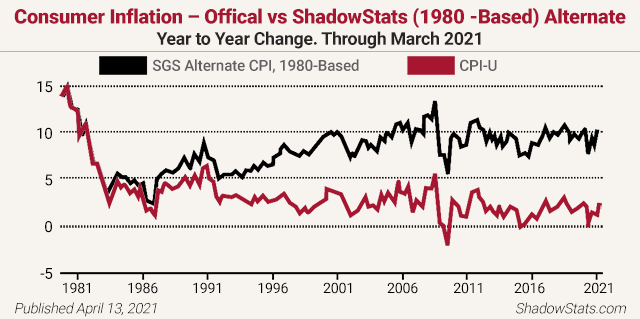
Analyst John Williams of the remarkable www.shadowstats.com website cites credible evidence of more than a 10% inflation rate instead of the government’s reported sub-2% rate. He notes, “The basic elements for a dollar collapse and an eventual hyperinflationary environment in the U.S. remain locked in place.” Williams reports inflation is over 11% right now (not the paltry 2% the Fed claims) and is about to take off, back above those pre-1981 levels you see below!
Even the normally bullish, pro-Wall Street editorial page of The Wall Street Journal conceded, “Fasten your seat belts and get ready for more
bumpy flying in the Fed’s cloudy skies.”
There is Good News IF You are in the Know — Here’s Why!
In addition to side-stepping fake paper asset and bogus investment landmines, this manual offers extremely useful guidance on how to vastly improve your financial situation —because it shows you how the ground has changed under your feet and the precise steps you must take to prepare for what is to come! You need the Ultimate Hyperinflation Defense Manual because it contains information the Wall Street media will NEVER share with you, such as:
- Where savvy investors are placing their assets to prevent them from being eroded by inflation and a declining U.S. dollar.
- How to avoid the coming nightmare millions of people on fixed incomes now face: Rising costs of everyday needs across the board and a smaller retirement nest egg capable of buying far less.
- Special Analysis: Who is killing the dollar, and how they are doing it… by crumbling the “Three Pillars” that have protected the U.S. economy.
- Are we being paid off for our silence on hyperinflation?
- Predictions from the renowned financial analyst who said, “I am 100% sure that the U.S. will go into hyperinflation.”
- No, you don’t have to liquidate your assets to buy the protection of gold! Here’s how to act, where to buy gold, how to protect it and more!
- How to use the little-known “bypass trust” — an incredible tool even if you fall below the estate tax threshold of wealth — and why ignorance about it can be downright dangerous.
- Nine GREAT ways your family can help you shield your assets and protect them from hyperinflation AND the government.
- The “Rule of 72” that savvy investors use to know what their money is really worth.
- Three steps you must take right away to prepare for a possible banking collapse.
- Shocking details of America’s true financial situation the government is trying to hide from Wall Street investors — you need to know this while it’s very limited knowledge.
- Simple steps to TRUE diversification in an inflationary environment — using all new financial instruments not available to investors during the 1970s.
- The REAL reason your savings aren’t safe (is the government coming for your money?) and what you can do about it starting today!
- Easy steps to take so you can begin living outside the system and preparing for the coming collapse.
- Learn the critical gold ratio signaling an uptrend that the big boys use — and you too can get rich.
- The shocking truth about the dollar — our fake promissory notes — that the Fed probably doesn’t want you to know but it’s knowledge you must have.
- How smart investors are quietly “hedging” like the pros to beat the money creators at their own game.
- Get the inside skinny on what coin dealers and precious metals merchants do not want you to know about their “collectibles.”
- The 10 best places to go offshore with your wealth so the government can’t get at it — and the flimsy reason they use to try and seize it!
- The 10 worst investing mistakes you can make that could cost you a fortune.
- How to use the secret onshore tax haven and wealth builder used by the privileged, corporations and the very wealthy.
- How to lower your taxes and get leverage against the government in retirement.
Isn’t this the kind of hands-on information you need to successfully navigate through the increasingly treacherous waters of your financial future? The Ultimate Hyperinflation Defense Manual is the ultimate protocol and guide… and a reference you can use for preserving your wealth for life. All the work and research in this book is worth the cover price of at least $89.95. But you won’t have to pay that…
You won’t find the Ultimate Hyperinflation Defense Manual in stores. This work is exclusively for my dedicated family of readers and is packed with usable, money-saving, problem-avoiding advice. If you are currently invested in the stock market, are looking to protect your wealth, are planning your retirement or are already retired, then you need this manual right away!
My Risk-Free Offer to You: Get This Vital Manual Now and Save Money!
Let me send you a fully-guaranteed copy of the updated, revised, improved and expanded Ultimate Hyperinflation Defense Manual absolutely FREE. Just subscribe to The Bob Livingston Letter® and I will send you this manual that can save you tens of thousands of dollars and make for a worry-free ocean of surety and safety in these inflationary times ahead.
Here are TWO MORE REASONS for You to Act QUICKLY!
For everyone who subscribes, I will include two FREE BONUSES — hot off the presses! The first brand-new report is called Investing Rules To Live By: Templeton, Munger, Graham, Buffett, Kennedy and the ‘Seven Secret Principles of the Rich.’ In it you’ll discover:

- Five ways to profit even if the investing rules are rigged!
- Who you have to know best, what you need to not believe, why you need to do research and how to assess real risk and reward.
- Templeton’s Rules: What kind of returns do you expect? Should you speculate? What if you make a mistake?
- How the most successful investors make their millions (not in bankster-ese language, but in plain words, with reflection on how to apply these concepts to your own investing).
- Yes, you too can make a huge fortune by a specific kind of knowledge with the ONE investing habit you MUST live by and by never shying away from this most powerful predilection.
- The “dumb money” buys on emotion? What do the rich buy on? If you want to be rich, you need to know how they think.
- And more!
Investing Rules To Live By is a MUST HAVE for every reader of The Bob Livingston Letter®. This report retails for $39.95… but it’s FREE with this offer only. Don’t delay. The inventory we’ve planned for will go quickly. But I also promised one more Special Report…

The second brand-new report is called How to Use the Wealth-Preserving Secret of Sam Walton. In it you’ll discover how Walton learned lessons in business succession planning and resolved to use the simple planning concept for their own family — and you can too — to shield their wealth from thieving bureaucratic fingers and greedy government tax-andspenders. This is essential knowledge to plan for your family — and shows you how to keep the government’s hands off your family’s wealth for future generations. A $19.95 value, yours FREE!
A WARNING: Ignorance of just one of these tools can be downright dangerous. But knowledge has the potential to keep your assets growing and secure from any upheaval. That’s why I hope you’ll…
Get Started Today and We’ll Take All the Risk
To receive a FREE copy my Ultimate Hyperinflation Defense Manual and the two FREE Bonus Gifts, all I ask is for you to subscribe to The Bob Livingston Letter® for two years (24 issues) for just $69. That’s just a drop in the bucket compared to what you’ll save when you receive your copy of my book and the two FREE Bonus Gifts.
For only $69… less than 1/25th of the of the price of a single ounce of gold… you’ll have in your hands THREE powerful tools that will let you beat the rigged money game in your favor. But here’s the best part…
I am so confident your financial well-being will improve dramatically over the next 24-plus months, if you ever decide you are not satisfied with your subscription, we’ll return every penny you paid — even up to the last issue. And you can keep your FREE Gifts.
All you have to do is make your selection from the No-Risk FREE Gifts Subscription Certificate located below to subscribe to The Bob Livingston Letter® and we’ll rush your FREE copy of Ultimate Hyperinflation Defense Manual and your two FREE Bonus Gifts right away. That’s $198.85 in FREE Gifts and Savings!
Of course, you can’t put a price on protecting yourself and your loved ones against the painful times ahead… so please order today.
To your health and wealth,

Bob Livingston
P.S. I’ve never felt more urgency about protecting you and your wealth! And I truly believe my Ultimate Hyperinflation Defense Manual could be the single most important tool you’ll ever discover. Don’t let this book collect dust. Join The Bob Livingston Letter® for $69 today and receive the Ultimate Hyperinflation Defense Manual for FREE and two FREE Bonus Gifts. That’s $198.85 in FREE Gifts and Savings! Click Here today!
NO-RISK FREE GIFTS SUBSCRIPTION CERTIFICATE
 YES! Please include me as a member of The Bob Livingston Letter® family as indicated below:
YES! Please include me as a member of The Bob Livingston Letter® family as indicated below:
Best Value: — $69: Please subscribe me to The Bob Livingston Letter® for TWO years (24 issues) for only $69 (a $49 savings). Plus send all three FREE Gifts listed below:
- FREE Gift #1 ($89.95 value): Ultimate Hyperinflation Defense Manual
- FREE Gift #2 ($39.95 value): Investing Rules To Live By: Templeton, Munger, Graham, Buffett, Kennedy and the ‘Seven Secret Principles of the Rich’
- FREE Gift #3 ($19.95 value): How to Use the Wealth-Preserving Secret of Sam Walton


$69 + FREE S&H
That’s a total savings value of $198.85!
Book and Reports only — $149.85 $89 plus $8.95 shipping and handling ($97.95 total).


$89 + $8.95 S&H